Finding a treatment for infectious diseases requires a thorough understanding of the organisms that cause them. Analyzing the structures of the COVID-19 virus helps find vulnerable sites, which are potential targets for drugs and vaccines. But discovering drug molecules that work on these sites requires pharmaceutical researchers to sift through a virtual library of more than a billion molecules. Then they have to test multiple permutations until they find one that completely binds to the target, inhibiting the protease and stopping the virus from replicating.
Where Oracle comes in
Such research involves running compute-intensive workloads, including detailed simulations of realistic biological systems that contain hundreds of thousands of atoms (Figure 1). In most on-premises clusters, complex simulations are typically queued based on resources available. However, testing many potential drug molecules, docking onto their targets through molecular simulations is often too compute-intensive for many on-premises clusters, which can take up to several months to produce results.
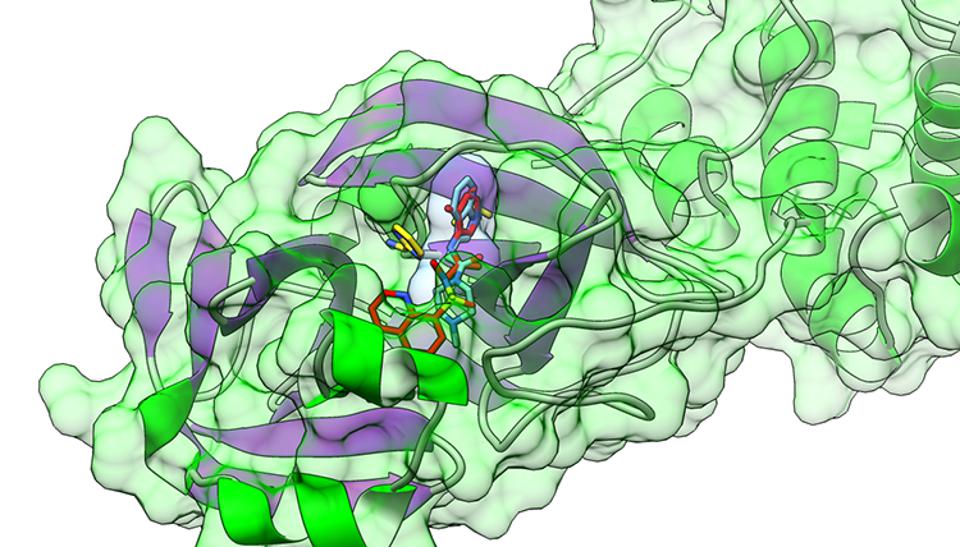
Figure 1: Rendering of the COVID-19 main protease, with the protein backbone in solid colors, and the protein surface shown partially transparent. The multi-colored sticks-figure depicts a drug molecule, which is binding to the active site of the protease. (Courtesy: Andy Jennings Consulting)
In a new “Oracle HPC in Healthcare, Technology Edition” webcast, Taylor Newill, director of high-performance computing (HPC) at Oracle, Mark Ross, cofounder of cloud-based rendering platform, GridMarkets, and Andrew Jennings, a computational chemist, joined me to explore how Oracle Cloud Infrastructure (OCI) is helping to solve this challenge by harnessing the power of HPC.
Simple and flexible, at scale
To help speed the process of running complex molecular simulations, such as the ones used to discover new treatments for COVID-19, GridMarkets runs its platform on OCI. Simulations are automatically provisioned with any number of Compute resources (bare metal and virtual machines) on demand. Whether simulating a drug molecule of 20 atoms with quantum mechanics to learn how electrons behave, or assessing multiple molecules made up of 2 million atoms, these tasks can take weeks using a cluster of traditional on-premises high-performance computers. Throw in simulations of drug molecules binding with different proteins, and it can take several months.
“These things require enormous calculations,” Jennings says. “Not even big pharmaceutical companies can justify buying an on-premises cluster big enough to speed through a few bursts, because for the better part of the year, that cluster just sits idle.”
Since Jennings began using GridMarkets to run molecular simulations in 2018, he has run complex molecular models in less than 24 hours.
Built for speed
With Oracle’s large-scale, high-performance cloud infrastructure, drug researchers don’t need to wait weeks to run their simulations or to scale down their biological systems to accommodate limited computing resources. Using molecular operating environment software on GridMarkets and OCI HPC, “we are not compromising, that’s the important thing here,” Jennings emphasized. “This is the opportunity where we can do exactly what we want, as quickly as we want. And that’s unprecedented.”
One of the things Jennings likes most about GridMarkets is that he can select how many machines he wants to run his simulations on, and then hit go. In a matter of seconds, GridMarkets configures the software and Compute resources and encrypts the data. When the job finishes, the machine shuts down, so there are no lingering costs. “None of this ties up local resources, I don’t have to sit behind a company’s firewall, and I can do it from home on a laptop,” he says.
Custom configurations, standard low cost
Because not all simulations used for early-stage drug discovery are programmed similarly, most require different resource configurations (Figure 2). Some simulation codes, such as molecular docking codes, scale well on a single instance with large number of CPUs or on an HPC cluster with multiple nodes. Other types of workloads, such as molecular dynamics simulations, require GPUs to process billions of atomic and molecular models.
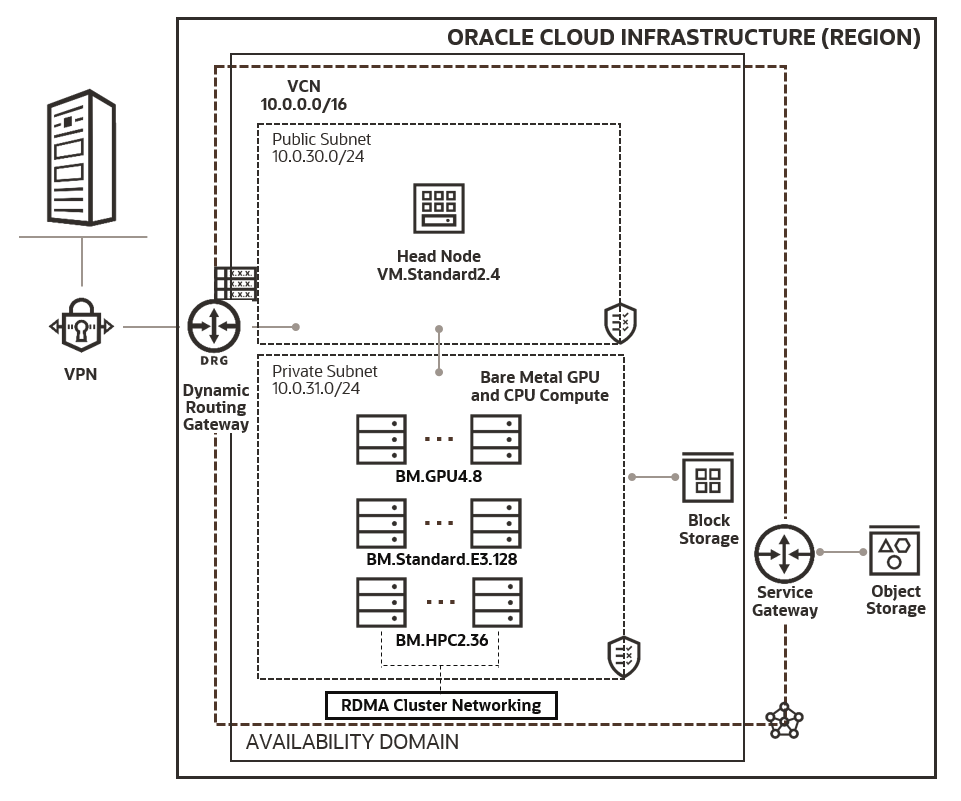
Figure 2: A sample architecture diagram of a large-scale cloud infrastructure for running simulation codes on multiple resource configurations. Resources inside the Private Subnet can be provisioned and destroyed on-demand to meet the different scale requirement.
Since the GridMarkets platform runs on high-performance servers located in Oracle Cloud data centers around the world, drug researchers like Jennings can access an unlimited number of resources whenever they need them, without having to pay for unused capacity when they don’t. Such pay-for-use capacity is also how GridMarkets keeps its own costs low. “We don’t own or maintain any hardware, and we’re paying 70% less to deploy an instance on Oracle Cloud Infrastructure than we did running workloads on Amazon Web Services or Google Cloud,” said GridMarkets co-founder and CEO Mark Ross.
OCI has made the latest CPU and GPU hardware (Figure 3) available to enable drug researchers, like Jennings, run their simulations on the most suitable hardware for their specific workloads.

Figure 3: Oracle Cloud Infrastructure offers the fastest and the beefiest bare metal CPU and GPU instances in the public cloud.
Conclusion
OCI HPC clusters feature instances with the fastest CPUs available in the public cloud. These instances are connected over dedicated RDMA cluster networking with 100-Gbps bandwidth, and less than 2-microseconds latency. OCI has also made the latest NVIDIA A100 GPUs available to enable the best performance for molecular dynamics simulations running on GPUs. Along with AMD EPYC-based E3 bare metal instances with 128 physical CPU cores and 2 TB of RAM, OCI provides state-of-the-art HPC infrastructure in the public cloud. As a result, Jennings and researchers like him using OCI can achieve the best possible computational performance, and reduce the time for discovering new treatments.
To know more about Oracle Cloud Infrastructure HPC roadmap, see Oracle Cloud Infrastructure: Compute and High-Performance Computing Roadmap Update. To experience the best HPC performance on Oracle Cloud, register for a free trial.